INTRODUCTION
According to government data in Colombia, Boyacá is the fifth largest milk producer, providing more than 1 184 000 litres daily. The production comes mainly from specialized dairy farms, which use genetic improvement, mechanized milking, milking routines, and correct storage of milk until its commercialization, however, mastitis is still a major concern for farmers due to the harmful effects on bovine health and the economic performance of farms, which for Colombia was calculated at up to US$ 25 000 dollars per farm (Ortíz-Duran et al., 2017; SIC, 2021).
Coagulase-negative Staphylococci (CNS) can be found on the skin of the udder and teats, as well as inside the teat canal, and are also known as environmental staphylococci. Despite being considered minor pathogens, they are the main bacteria associated with cases of subclinical mastitis when the major pathogens are controlled (Dufour et al., 2012; Krishnamoorthy et al., 2016, Levinson et al., 2016, Andrade-Becerra et al., 2021; Ruiz-Romero & Vargas-Bello-Pérez, 2023). Also, cases of the presence of genes associated with antibacterial resistance in CNS isolated from bovine subclinical mastitis have been reported in multiple countries (Jiménez et al., 2020, Crespi et al., 2022, Ibrahim et al., 2022). This is especially important because, being reservoirs of these genes, they can transmit them horizontally to other bacteria of the same genus (Petersen et al., 2013). Antimicrobial resistance is a serious concern, mainly due to the implications for public health, and the economic costs associated with parenteral or intramammary treatment failures, milk, and animal losses (Crespi et al., 2022). This problem is related to mistakes in pharmacological prescription, misdiagnosis of the etiological agent, and failures in the development of treatment (Petersen et al., 2013; Crespi et al., 2022, Ibrahim et al., 2022).
The presence of the mecA and mecC genes in methicillin-resistant staphylococci (MR-staphylococci) has been associated with resistance to β-lactams, as well as to groups such as aminoglycosides, tetracyclines, macrolides, chloramphenicol, and fluoroquinolones (Baig et al., 2018; Ibrahim et al., 2022). The source of this resistance is that the gene translates into a penicillin-binding protein (PBP) that has a low affinity for the medicament, thus preventing its action. This gene is in a mobile element of the bacterial genome called the staphylococcal chromosomal cassette (SCCmec), which has been divided into 13 types (I-XIII) and multiple subtypes due to genetic and structural changes of the protein (Lakhundi & Zhang, 2018; Crespi et al., 2022).
Epidemiological analyses of the etiology of mastitis are important because they allow the identification of risk factors present on farms, as well as the most common pathogens and their possible effects on milk quality. Previous reports done by the author’s group have shown the presence of CNS as causal agents of bovine subclinical mastitis (Tarazona-Manrique et al., 2019) and its increasing effect on the somatic cell count in milk (Andrade-Becerra et al., 2018. 2021). In addition, other reports in the country show the importance of these agents as causes of subclinical mastitis in dairyproduction systems (Calderón et al., 2011; Sánchez &Gutiérrez, 2015; Jiménez et al., 2020). Therefore, this study aimed to determine the antibiotic resistance profile and the presence of the mecA gene in antibiotic resistance CNS species isolated from subclinical mastitis of Holstein cows in Boyacá.
MATERIALS AND METHODS
Coagulase-Negative Staphylococci Species
The CNS species used in this study come from a local study (Andrade-Becerra et al., 2021), in which 619 samples obtained over 6 months in one dairy farm were positive for at least one CNS species. The most prevalent species were S. chromogenes (18.6%), S. epidermidis (14.0%), S. simulans (12.3%), S. capitis (11.3%), S. sciuri (10.5%), S. haemolyticus (9.4%), S. hyicus (5.8%), S. hominis (5.5%), S. auricularis (5.2%), S. xylosus (3.7%), S. equorum (1.3%) and the 2.4% of the samples were positive to multiple CNS species. Bacterial DNA was extracted with the AllPrep Bacterial/Fungal DNA/RNA/Protein kit (Qiagen, Germany) following the manufacturer’s instructions, and stored at -80 °C until use in the present study.
Antibiogram Profile of CNS Species
At the time of bacterial isolation from samples of cows with subclinical mastitis, all isolated CNS were subjected to an antibiogram on Müeller-Hinton agar (Merck KGaA, Germany) using the following antibiotics: Ampicillin 10 µg, Tetracycline 30 µg, Streptomycin 15 µg, Oxacillin 1 µg, Penicillin 10 µg, Enrofloxacin 5 µg (Becton Dickinson, USA). The strains were classified as susceptible, intermediate, or resistant according to the size of the inhibition halo after incubation at 37 °C for 24 h and comparing it with the manufacturer’s instructions for the sensi-discs.
Gene mecA Detection in CNS
The detection of the mecA gene was performed as indicated by Ibrahim et al. (2022). T he PCR reaction was performed using SIMPLIAMP (Applied Biosystem) endpoint thermocycler in a final volume of 25 µL. The primers used for mecA gene detection were: Forward: GTA GAA ATG ACT GAA CGT CCG ATA A, and Reverse: CCA ATT CCA CAT TGT TTC GGT CTA A, with an expected amplicon size of 310 bp (Spanu et al., 2004). The protocol for the amplification cycles had initial denaturation at 95 °C for 4 min, followed by 35 cycles of denaturation at 95 °C for 30 s, annealing at 58 °C for 40 s, and extension at 72 °C for 1 min, with a final extension at 72 °C for 10 min. The result was observed on a 1.5% agarose gel. Two negative controls with 2 µL of sterile water were used instead of the sample. Positive controls were not used because they were not available (Ibrahim et al., 2022).
RESULTS
Of all the CNS isolates, S. hyicus, S. hominis, S. auricularis, S. xylosus y S. equorum did not show resistance to any of the antibiotics evaluated, corresponding to 20.46% of the isolates (127/619). In addition, the samples that showed mixed growth were not included (2.42%, 15/619). The other CNS species have full or intermediate resistance to the six antibiotics evaluated (Table 1).
Table 1. Number of isolates of CNS from bovine subclinical mastitis resistant (intermediate or completely) to at least one antibiotic
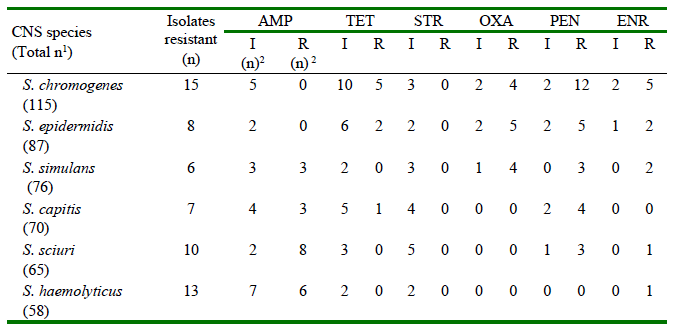
1 Total number of isolates obtained for each of the CNS species
2 I(n)/R(n) Number of isolates that had intermediate or complete resistance to the drug evaluated, respectively, according to the diffusion test
AMP: ampicillin 10 μg; CEF: ceftriaxone 30 μg; TET: tetracycline 30 μg; STR: streptomycin 15 μg; OXA: oxacillin 1 μg; PEN: penicillin 10 μg; ENR: enrofloxacin 5 μg
Based on results shown in Table 1, percentages of isolates phenotypically resistant to at least one antibiotic used in the agar diffusion test were found: 13.04% of S. chromogenes, 9.19% of S. epidermidis, 7.89% of S. simulans, 10% of S. capitis, 15.38% of S. sciuri, and 22.41% of S. haemolyticus. Percentages of full and intermediate resistance determined for each of the antibiotics evaluated are shown in Figure 1.
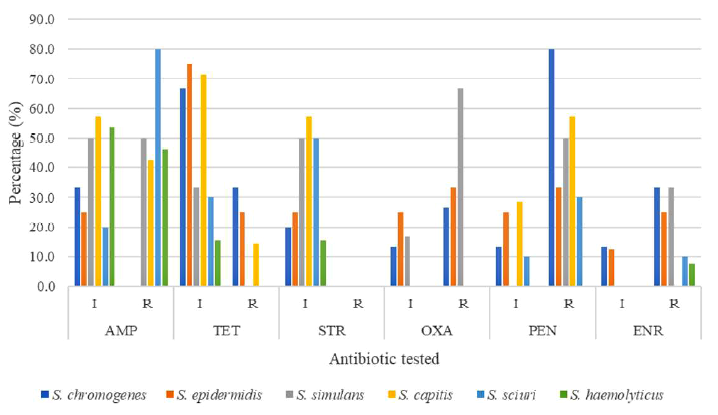
Figure 1. Percentage of Intermediate or full Resistance in Coagulase-negative Staphylococci species (CNS) isolates per antibiotic tested in agar diffusion test
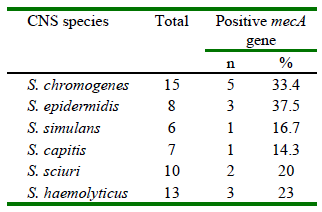
Based on the above results, CNS species that were resistant to at least one antibiotic were subjected to molecular analysis for the detection of the mecA gene through endpoint PCR using specific primers. The results can be seen in Table 2. Interestingly, more than 14% of the isolates resistant to drugs were positive for the presence of the mecA gene.
DISCUSSION
All isolates showed intermediate resistance to ampicillin, tetracycline, and streptomycin, however, not all had complete resistance to all antibiotics tested, which indicates that antibiotic resistance patterns vary between CNS species even when they belong to the same farm (Figure 1). Also, it is interesting to note that some species like S. capitis, S. sciuri, and S. haemolyticus have the mecA gene but they do not show resistance to both oxacillin and penicillin. This indicated that the presence of the gene by itself is not necessarily related to the resistance to the antibiotics.Also, it shows that the mecA gene activation can be influenced by other factors like other resistance-related genes or the mammary gland environment.
Studies in Colombia have revealed the importance of CNS as the cause of bovine subclinical mastitis. Jiménez et al. (2020) performed a study in four Colombian departments (none of them were Boyacá) and determined that the isolated CNS species were commonly more resistant to ampicillin and penicillin, associating it with the presence of the blaZ gene. Despite the similarity of species and antimicrobial resistance present between the two studies, which would indicate the importance of these as causal agents of mastitis in national dairies, it also indicates similarity between livestock practices, especially in terms of antibiotic administration for treatment. It is important to note that in this study (and contrary to what happened in Jiménez et al., 2020) the determined CNS species had marked resistance to tetracycline and streptomycin, which is related to Sánchez & Gutiérrez (2015) study in Tolima, who determined that 58% of the CNS samples were resistant to tetracyclines.
Jiménez et al. (2020) also determined the presence of the mecA gene in methicillinresistant CNS isolated from bovine mastitis milk in 27% of the samples evaluated, contrasting with the results obtained in a previous study (Calderón et al., 2011). The current results show the presence of the mecA gene in up to 37% of the samples of S. epidermidis resistant to multiple antibiotics through the agar diffusion test, evidencing that the presence of the gene varies even between CNS species even though they are isolated from the same farm, which is a finding not previously reported in Colombia. This shows that in little more than a decade, reports of multiple antimicrobial resistance have grown, and, therefore, epidemiological investigations in this regard must increase to make better decisions for prevention, control, and treatment of the disease caused by CNS.
Latin American studies such as Crespi et al. (2022) showed that 24.6% of CNS isolates from bovine mastitis were resistant to penicillin, followed by erythromycin, clindamycin, cefoxitin, and oxacillin, results like those found in this study. In the case of S. chromogenes, S. epidermidis, and S. simulans, the results are corroborated by the study conducted by Raspanti et al. (2016). Likewise, Gentilini et al. (2002) showed higher CNS resistance patterns mainly to penicillin. However, Soares et al. (2012) differ slightly from this and other studies as they reported S. xylosus, S. cohnii, S. hominis, S. capitis, and S. haemolyticus as the main CNS in Brazil, nevertheless, the antimicrobial resistance profile and the association of the blaZ and mecA genes were identical to what was reported in this study.
The above would indicate that, regardless of the type of CNS present as the causative agent of mastitis, all can carry resistance genes and transmit them to other CNS species; however, those bacteria that have better multiplication and maintenance mechanisms in the medium will be more likely to be transmitted within the herd, which is why analysis of risk factors is an important aspect in determining these epidemiological differences. For example, some species, such as S. chromogenes and S. haemolyticus may have colonization advantages in the mammary gland because of hemolytic and proteolytic activity, better tolerance to postmilking teat disinfection, and the capacity to elude the immune system (Hosseinzadeh & Dastmalchi, 2014). These bacteria have the potential to spread zoonotic diseases and provide antibiotic-resistance genes to humans who interact with dairy cows, and vice versa (Fergestad et al., 2021).
Sawant et al. (2009) showed that most CNS isolates were susceptible to several antibiotics with the exceptions of S. chromogenes, S. epidermidis, and S. hyicus, which showed phenotypic resistance to penicillin, either through constitutive or inducible production of β-lactamase, associating it with the presence of the blaZ gene. In addition, they determined that 32% of the isolated S. epidermidis were positive for the presence of the mecA gene, which agrees with results obtained in the present study and those reported by Taponen et al. (2016). Furthermore, Sawant et al. (2009) showed that all multi-drug-resistant S. epidermidis were genotypically related despite being isolated from different cows on the same farm suggesting clonal spread, and this may be happening on the farm evaluated because all the isolates come from different cows at different times of lactation and in different months.
The presence of multiple CNS species causing subclinical mastitis in a herd and the diversity of both phenotypic and genotypic drug resistance patterns shown by the different isolates show the importance of applying microbiological culture and susceptibility testing, in addition to detection resistance genes through PCR should be performed routinely on farms to develop disease control and treatment strategies more effectively.
Regarding antimicrobial resistance profiles in Egypt and South Africa, Ibrahim et al. (2022) & Phophi et al. (2019) determined a high frequency of CNS isolates were resistant to penicillin and oxacillin, followed by resistance to clindamycin, chloramphenicol, amoxicillin, and erythromycin; in addition, all the isolates were resistant to several antibiotics at the same time. The results found in this study show a similar dynamic, where all the isolates present resistance to some degree to several antibiotics at the same time, also determining that S. chromogenes and S. epidermidis had intermediate antibiotic resistance to all the drugs evaluated and complete resistance to TET, OXA, PEN, and ENR.
In conclusion, the results show differences in the pattern of resistance to multiple antibiotics in the CNS species, even though they were isolated from the same farm for six months. The presence of the mecA gene is an important finding because it corroborates previous findings in the country and demands increasing research related to determining its presence, as well as other genes that may be related to resistance to multiple antibiotics that were found, since the presence of a single gene cannot be associated with this wide range of resistance to multiple antibiotics. It is also necessary to determine the risk factors associated with the presence of these CNS species on farms and to determine if there is a bacterial interchange between milkers and cattle, due to the possible public health implications.











 uBio
uBio