Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO
Related links
-
 Similars in
SciELO
Similars in
SciELO
Share
Revista Peruana de Medicina Experimental y Salud Publica
Print version ISSN 1726-4634
Rev. perú. med. exp. salud publica vol.37 no.4 Lima Oct-Dec 2020 Epub Nov 18, 2020
http://dx.doi.org/10.17843/rpmesp.2020.374.5169
Original articles
Infectious agents in biological samples from patients with Guillain-Barré syndrome in Peru, 2018-2019
INTRODUCTION
Guillain-Barré syndrome (GBS) is an acute polyneuropathy that manifests as an acute ascending flaccid paralysis. Its diagnosis is mainly clinical, based on the anamnesis and findings during physical examination, complemented by electromyographic results and the typical changes observed in the cerebrospinal fluid (CSF) 1. The physiopathology of GBS is based on an immunological phenomenon, generally triggered by an infectious process. It has been postulated that this immune response is initiated by a wide variety of antigens, such as infections by virus or bacteria, particularly Campylobacter jejuni 2. A systematic review has estimated that between 40 and 70% of the cases of GBS are preceded by an upper respiratory infection or an infection of the gastrointestinal tract; mainly due to C. jejuni 3 which is the cause of GBS in up to 31% of cases 2. In a meta-analysis, inactivated monovalent influenza A (H1N1) virus vaccine was reported to increase the risk of GBS by 1.6 cases per million doses applied, a small increase 4.
Zika is one of the viruses reported as possible etiological agents of GBS. Outbreaks of Zika associated with GBS have been reported studies carried out in French Polynesia 5, Brazil, Colombia 6, Dominican Republic, El Salvador, Honduras, Suriname, and Venezuela 7. A bias in all these studies is that no tests have been conducted to rule out etiological agents other than Zika, dengue, and chikungunya.
Viruses from the genus Enterovirus are associated with acute flaccid paralysis (AFP), with a clinical picture like GBS. This genus has 12 species including enteroviruses A to J and rhinoviruses A, B and C. The transmission route depends on the serotype, some are transmitted by the fecal-oral route and others by the respiratory tract. Many enteroviruses are neurotropic and can affect the central or peripheral nervous system. For example, Coxsackievirus and echovirus can cause viral meningitis; Coxsackie, enterovirus A71 and echovirus 9 can produce meningoencephalitis 8.
Poliovirus and enteroviruses D68 and A71 can produce AFP. In 2014, an increase in acute respiratory infection caused by EV-D68 was observed in the United States as well as an AFP outbreak associated with EV-68 9. Other viruses, such as Coxsackie and echovirus, were reported to cause AFP in Korea, with a low incidence 10. GBS incidence varies with age, seasonality, and the place where measurements are made. Considering all age groups, the incidence varies from 0.16 to 3 × 105 person-years, is higher in men and increases with age 3. Peru has had an unusual increase of GBS cases since 2018, with 1,378 cases reported and an incidence of 4.59 × 105 inhabitants, compared to previous years, which was approximately 0.91 × 105 inhabitants 11. In 2019, until the epidemiological week number 47, a total of 1,341 cases were reported 12. A high percentage of the cases occurred in the northern and central departments of the country (Lambayeque, La Libertad, Piura, and Lima). However, almost all of Peru’s departments were affected by the outbreak. Several hypotheses have been proposed regarding possible infectious agents associated with GBS 13; however, information about affected patients and the results of the tests conducted have not been systematized.
The aim of this study is to describe the results of laboratory tests performed on biological samples from patients diagnosed with GBS received at Instituto Nacional de Salud between 2018 and 2019.
KEY MESSAGES
Motivation for the study: Due to the unusual increase in cases of Guillain-Barré syndrome (GBS) between 2018 and 2019, we conducted a search for the possible etiologic agent that triggers the autoimmune process that causes this neurological syndrome.
Main findings: We identified several pathogens, among which we observed a variant of Campylobacter jejuni associated with GBS outbreaks in other countries. We propose this pathogen as a probable causal agent of the outbreaks in Peru.
Implications: These findings open the opportunity to identify the infectious agent that triggers GBS and will help to implement prevention and control measures to cut the chain of infection of Campylobacter jejuni.
MATERIALS AND METHODS
Type of study and population
We carried out an observational and descriptive study on patients with GBS treated in the hospital networks of Ministerio de Salud (MINSA), Seguro Social (EsSalud) and Sanidad de las Fuerzas Armadas y Policiales, from which biological samples were obtained to be analyzed in the Instituto Nacional de Salud (INS) or processed in the Regional Reference Laboratories. To contextualize the outbreak, we reviewed the reports on the website of the Centro Nacional de Epidemiología, Prevención y Control de Enfermedades (CDC-Peru) on the number of GBS cases reported per epidemiological week since the beginning of 2018.
The INS is the state agency in charge of public health research and specialized diagnosis. INS and CDC-Peru work on the investigation of unusual health occurrences and epidemic outbreaks. As part of the investigation of an outbreak, immunological, chemical, phenotypic and molecular laboratory tests, among others, are performed to explore the association with infectious agents, chemicals, or both.
We included patients from the epidemiological surveillance system who sent biological samples to the Regional Reference Laboratories of Lima, Piura, Lambayeque, La Libertad, Cajamarca, Junín or to the INS to rule out infectious diseases. We also described the report of a private laboratory that processed biological samples from eight patients and sent the results to the INS.
In Peru, since 2016, the GBS has been a notifiable disease for the Epidemiological Surveillance System, in all public and private health facilities, according to the emergency health protocol for GBS surveillance 14. This standard defines a clinically confirmed case of GBS (level 3 of diagnostic certainty, according to the Brighton criteria) when the following criteria are met: bilateral flaccid muscle weakness in the extremities; diminished or absent deep osteotendinous reflexes in the weakened extremities; monophasic disease, with an interval between the onset and nadir of the weakness between 12 hours and 28 days, followed by a clinical plateau; and absence of alternative diagnosis to explain the weakness. GBS may or may not have electrophysiological findings in the CSF regarding albumin-cytological dissociation. In June 2019, CDC-Peru, in response to the increase in GBS cases in northern Peru, issued an epidemiological alert notifying of all clinical cases of GBS along with the recollection of the following biological samples: serum, feces, nasopharyngeal swabs, CSF, and urine 15.
Procedure
Serum tests used were IgM and IgG ELISA for Dengue, Zika and Chikungunya viruses; and quantitative polymerase chain reaction (qPCR) for Dengue and Zika viruses. IgM ELISA and polymerase chain reaction (PCR) in CSF were used for Dengue and Zika viruses; PCR for enterovirus, as well as the FilmArray® meningoencephalitis/encephalitis panel. We performed real-time reverse transcription polymerase chain reaction (qRT-PCR) on nasopharyngeal swab samples to detect influenza A and B viruses, rhinovirus, respiratory syncytial virus, and adenovirus. We used direct immunofluorescence for influenza A and B viruses, respiratory syncytial virus, adenovirus, parainfluenza virus 1, parainfluenza virus 2, parainfluenza virus 3, and metapneumovirus. We used rectal swabs and stool cultures to search for enteropathogenic bacteria and enteroviruses, as well as PCR for enteroviruses, Campylobacter spp., diarrheagenic Escherichia coli and Shigella spp.
Any PCR result that indicated the presence of genetic material from any of the mentioned microorganisms was considered positive, as well as the predetermined values of ELISA results for each etiological agent 16. In the case of results submitted to the INS by other laboratories, the method mentioned in the report was shown.
We selected the infectious agents according to their frequency to be subjected to sequencing of the entire genome. In the case of viruses, the sequencing was done with the Sanger method 17. The sequencing result was obtained in the ABI format, and the sequences were assembled and analyzed using the Geneious V8 software. Likewise, the Blast and Kallisto algorithms were used to detect the most similar transcripts 18.
Four CSF samples and three stool samples collected in 2018, which tested positive for enterovirus at the INS, were sent to the Center of Diseases Control of the United States of America (CDC-USA), where PCR was performed for enterovirus and human parechovirus.
Statistical analysis
We obtained the laboratory results from the NetLab application, the INS laboratory results reporting system, and exported them to an Excel database (Microsoft Office®). Categorical variables were described using relative and absolute frequencies. For the numerical variables, their distribution was determined by means of the Shapiro-Wilk test and they were described in means and standard deviations or medians and interquartile range, respectively, if they fulfilled or not the normality criterion. The analysis was performed in Stata version 9 (StataCorp, College Station, Texas, USA).
Ethical aspects
This study is part of the investigation of the GBS outbreak in Peru. NetLab registry codes were used without identifying the patients. Laboratory results were only used in the context of this epidemiological situation, so it was not necessary to submit them to an institutional ethics committee for approval.
RESULTS
The INS received a total of 2,051 biological samples including serum (738), CSF (247), nasopharyngeal swab (475), urine (278), rectal swab and/or stool (313). These samples were obtained from a total of 906 patients with a clinical diagnosis of GBS from April 27, 2018 to November 5, 2019, of which 26 (2.9%) samples were obtained in 2018 and 880 (97.1%) in 2019. Figure 1 shows the number of patients with GBS reported by the CDC-Peru and the number of patients whose biological samples have been sent to the INS (included in this study) in the mentioned period.
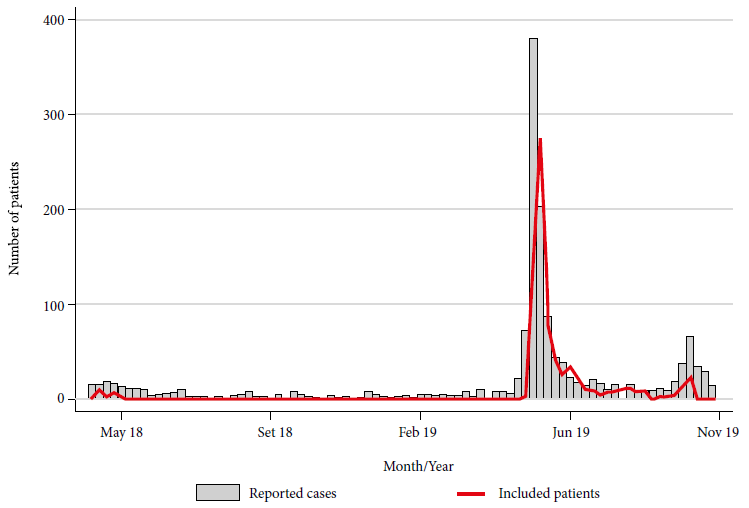
Figure 1 Cases of Guillain-Barré syndrome reported by Centro Nacional de Epidemiología, Prevención y Control de Enfermedades and patients with biological samples sent to Instituto Nacional de Salud, Peru, 2018-2019.
Most patients were from Lima, Piura, La Libertad, and Junín (Table 1), and from the following hospitals: 52 (5.9%) patients were from the Hospital de Apoyo Santa Rosa, Piura; 49 (5.6%) from Hospital Nacional Almanzor Aguinaga Asenjo, Chiclayo; 48 (5.5%) from Hospital Edgardo Rebagliati Martins, Lima; and 43 (4.9%) from Hospital Nacional Daniel Alcides Carrión, Junín.
Table 1 Demographic characteristics and provenance of Guillain-Barré syndrome cases with biological samples sent to Instituto Nacional de Salud, 2018-2019 (n = 906).
| General characteristics | n (%) |
|---|---|
| Median age (IQR) | 40 (24-56) |
| 0-14 | 112 (12.7) |
| 15-24 | 113 (12.8) |
| 25-40 | 207 (23.4) |
| 41-60 | 271 (30.7) |
| >60 | 181 (20.5) |
| No data | 22 (2.4) |
| Male sex | 539 (59.5) |
| Provenance | |
| Lima | 358 (39.5) |
| Piura | 199 (22.0) |
| La Libertad | 98 (10.8) |
| Junín | 82 (9.1) |
| Lambayeque | 61 (6.7) |
| Cajamarca | 48 (5.3) |
| Others | 60 (6.6) |
IQR: Interquartile Range
From the 738 serum samples, 735 had negative serology or PCR results for arbovirus; 3 patients were positive for dengue and came from Lambayeque, La Libertad, and Lima. Likewise, 3 patients had positive PCR in urine for Zika virus. Of the CSF samples, 12 were positive by PCR for enterovirus; however, all were from 2018 and all cultures were negative. Five of these patients also had positive PCR for enterovirus in their stool. Of the 220 CSF samples from 2019 analyzed by PCR and culture for enterovirus, none were positive; the 90 samples processed by the FilmArray technique were negative for the 14 pathogens identified by the test. Regarding respiratory viruses, 1 sample was positive for influenza A virus, 2 for influenza B virus, 2 for adenovirus, five for respiratory syncytial virus, and ten for rhinovirus. Finally, in 24 patients, stool/rectal swab culture was positive for Campylobacter spp. and 69 for diarrhoeagenic E. coli (Table 2).
Table 2 Laboratory results of biological samples sent to Instituto Nacional de Salud, from patients with Guillain-Barré syndrome, Peru, 2018-2019.
| Sample type | Diagnostic test | Positives n (%) | Total of processed samples |
|---|---|---|---|
| Serum | Dengue IgM ELISA | 2 (0.5) | 369 |
| qRT-PCR Dengue | 1 (1.9) | 54 | |
| qRT-PCR Zika | 0 (0) | 372 | |
| Zika IgM ELISA | 0 (0) | 597 | |
| Chikungunya IgM ELISA | 2 (0.3) | 581 | |
| qRT-PCR Chikungunya | 0 (0) | 15 | |
| Urine | qRT-PCR Zika | 3 (1.1) | 278 |
| CSF | qRT-PCR Zika | 0 (0) | 82 |
| IgM Zika ELISA | 0 (0) | 64 | |
| qRT-PCR Dengue | 0 (0) | 5 | |
| IgM Dengue ELISA | 0 (0) | 1 | |
| RT-PCR Enterovirus | 12 (5.1) | 234 | |
| Culture of Enterovirus | 0 (0) | 222 | |
| FilmArray® Meningitis/Encephalitis panel | 0 (0) | 90 | |
| Swab | DIF Influenza A virus | 0 (0) | 293 |
| qRT-PCR Influenza A virus | 1 (0.2) | 475 | |
| DIF Influenza B virus | 1 (0.3) | 304 | |
| qRT-PCR Influenza B virus | 1 (0.2) | 464 | |
| qRT-PCR Rhinovirus | 10 (5.6) | 179 | |
| qRT-PCR RSV | 5 (2.8) | 177 | |
| Dif RSV | 0 (0) | 171 | |
| qRT- PCR Adenovirus | 2 (1.1) | 176 | |
| DIF Adenovirus | 0 (0) | 171 | |
| DIF Parainfluenza 1 | 0 (0) | 171 | |
| DIF Parainfluenza 2 | 0 (0) | 171 | |
| DIF Parainfluenza 3 | 0 (0) | 171 | |
| DIF Metapneumovirus | 0 (0) | 171 | |
| Rectal swap | RT-PCR Enterovirus | 5 (2.0) | 250 |
| Campylobacter spp (by PCR and culture) | 24 (6.5) | 367 | |
| Diarrheagenic E. coli | 69 (22.9) | 302 |
CSF: cerebrospinal fluid, qRT-PCR: real-time quantitative reverse transcription polymerase chain reaction; RT-PCR: reverse transcription polymerase chain reaction; PCR: polymerase chain reaction; DIF: direct immunofluorescence.
A private laboratory in Lima reported to INS the results obtained by the FilmArray® gastrointestinal panel, from 8 patients with GBS reported in 2018. There were 5 positive results for enteropathogenic E. coli, 4 for Campylobacter spp., and 3 for human rhinovirus/enterovirus, Shigella spp., enteroinvasive E. coli, and enteroaggregative E. coli.
We carried out the molecular study of CSF and stool samples received by the INS with positive PCR results for enterovirus. First, the 5' UTR region was amplified and identified as enterovirus. Then, the complete genome was sequenced; however, the number of fragments detected was less than 10% of the total virus genome, so it was not possible to identify the species. Out of 4 CSF samples and 3 stool samples with positive PCR for enterovirus sent to the CDC-USA, only 1 stool sample was positive for echovirus 1.
Similarly, the complete genome of the 10 strains isolated from Campylobacter spp. was sequenced: 6 biological samples received at the INS (two from Piura and four from Junín) and 4 strains isolated by the Naval Medical Research Unit 6 (NAMRU-6). All specimens were from 2019. We proceeded according to the methods previously described and identified the C. jejuni species. The phylogenetic analysis of this strain’s genomic sequence identified it as genotype ST2993, a genotype never reported in Peru. The ten isolates are from the same clone, which means, they share the same genetic information. As part of the genomic analysis, we also identified the presence of cst-II and cst-III genes, which code for lipooligosaccharides associated with GBS. Likewise, we observed the presence of a specific point mutation in the quinolone resistance determining region (QRDR) of the gyrA gene (Thr86Ile), which is related to resistance to fluoroquinolones.
DISCUSSION
We were able to systematize the laboratory results of patients diagnosed with GBS in Peru whose biological samples were analyzed at the INS between 2018 and 2019, as well as those whose results were submitted to the INS. Medical literature reports that up to 75% of GBS cases have a history of previous respiratory or digestive infection in the last six weeks; however, in many of them, the infection had already been controlled by the time the etiological agent began to be searched, thus it couldn’t be isolated 19. The presence of C. jejuni (6.5%) stands out among all the infectious agents isolated or identified in this study. Other agents we identified are the following: arbovirus (1.1%), enterovirus (2%), diarrheagenic E. coli (22.9%) and respiratory viruses as rhinovirus in 5.6% and RSV in 2.8%.
This is the first report of isolation of C. jejuni ST2993 in Peru and America. A strain of C. jejuni of the same genotype ST2993 was reported in 2007 in China and was associated to a GBS outbreak 20. Previous reports of genetic sequencing of Peruvian isolates of Campylobacter show strains genetically quite different from this new strain 21. Ten isolated strains were genetically sequenced, and the results were indistinguishable from each other, so we conclude that it is the same clone with a wide distribution since it was identified in the regions of Piura, Junín, and Lima. There are multiple reports associating C. jejuni with GBS 2 , 20 , 21. Activated macrophages, T lymphocytes and serum antibodies against human gangliosides are related to the physiopathology of GBS and it is believed that the lipooligosaccharides of C. jejuni antigenically mimic these gangliosides, triggering the immune reaction 22. Therefore, this finding is significant and urges to investigate the chain of infection of C. jejuni, which is usually associated with poultry, dairy food consumption, water, meat, among others.
We identified diarrheagenic E. coli in stool/rectal swab samples from 69 patients. However, these strains that showed high heterogeneity corresponded to different pathotypes. There is little information on the association between this bacterium and the occurrence of GBS, and previous reports refer predominantly to urinary tract infections rather than enteric infections 23. We have not found reports of GBS outbreaks with this etiology, so the association with the outbreak in Peru seems unlikely, considering that E. coli is a germ that is part of the human gastrointestinal tract microbiome.
Regarding arboviruses, we identified infection by dengue virus in three patients (two by IgM antibodies and one by serum PCR) and infection by Zika virus also in three patients (positive PCR in urine). Additionally, two patients had positive IgM antibodies against chikungunya virus; one was from Piura and the other from Cajamarca (province of Jaén). There are several reports in the literature about the association between GBS and Zika virus infection 24. However, in our study there were only three cases which reflects the usual transmission in endemic areas of this virus. The same is true for chikungunya, an increase of GBS cases was reported during an outbreak of chikungunya in French Polynesia as well as other reports of sporadic cases 25. However, it should be considered that this outbreak of GBS has even affected areas of the central and northern highlands of Peru, where there is no transmission of these viruses. Recently, the role of Zika virus in these outbreaks has been questioned because most of the studies on this subject do not use a control group which does not allow to determine association 26.
Few patients test positive for respiratory viruses. Although the association between GBS and influenza virus is documented 27, we believe that due to the low frequency observed in this group of patients these viruses might not be relevant to the development of GBS in the outbreak in Peru.
The CDC-US laboratories (echovirus 1) and one Peruvian private laboratory identified enteroviruses in CSF of patients diagnosed with GBS in 2018, these results were consistent with the INS findings. However, all viral cultures for enteroviruses were negative, possibly because of an inadequate sample or some problem that may have affected the viability of the virus. Furthermore, this finding was limited to 2018. During the 2019 outbreak, no genetic material for enteroviruses was isolated or identified in any patient’s stool or CSF samples. There are reports in the literature associating enteroviruses with acute neurological syndrome, specifically enterovirus D68 9. However, these reports refer to acute flaccid paralysis with a neurological picture that affects the motor neuron located in the anterior horn of the spinal cord, a clinical and electrophysiological picture different from GBS. In the GBS the peripheral nerves are affected in each of the different forms of presentation 22, this has been reported in Peru. Based on these findings, we ruled out the epidemiological link of enterovirus with this GBS outbreak.
Papers on the study of the phylogeny of digestive pathogens in Peru are scarce and the few that exist have been conducted on children under five years old, which limits our knowledge about the microorganisms that circulate in our country. At a time when technological advances allow access to genome sequencing in public health, this knowledge gap must be quickly resolved with research or, ideally, with the implementation of specific sentinel surveillance.
There are some limitations to our study. It uses a descriptive design and does not have a control group, consequently, the association between GBS and a potential etiologic agent was not demonstrated; however, our hypothesis is that C. jejuni is the potential etiologic agent, which should be demonstrated with further analytical studies. The tests were carried out by regional laboratories and by the INS, under the usual parameters of quality control, without additional studies of concordance between laboratories. Only one stool sample or rectal swab was collected for stool culture to isolate Campylobacter spp., therefore the sensitivity of this test may be reduced. One study reported a sensitivity of 100% with three consecutive stool samples 28. In Peru, self-medication with antibiotics is relatively frequent, and can range from 34% in the general population 29 to 70% in university students 30; without this information about the patients, the sensitivity of stool tests could be reduced. The time between the collection of stool samples and the processing to carry out the stool culture in many cases was more than 24 hours, which reduces the sensitivity to isolate C. jejuni. The collection of biological samples was not carried out according to the surveillance system; samples such as CSF were collected only in 25.8%, urine in 30.7% and stool or rectal swab in 40.5% of the cases. Finally, serum samples were usually collected after the treatment had already started, which could alter the serological tests with false positive results in patients who were given immunoglobulins, or false negatives if plasmapheresis were used.
In conclusion, various pathogens were identified in biological samples from patients with GBS in the outbreaks that occurred in Peru between 2018 and 2019. The most outstanding finding from 2018 was the identification of enteroviruses by PCR at the INS; however, this finding could not be confirmed by genome sequencing and, in addition, of the four samples sent to the CDC-USA only one was positive for echovirus. All CSF and stool samples were negative for enteroviruses by culture and PCR in 2019. On the other hand, it was possible to isolate C. jejuni genotype ST2993 in 2019, a pathogen widely related to GBS in several continents. This finding opens new research lines to carry out analytical studies, identify the reservoir and the source of infection to implement the most effective prevention measures and cut the chain of infection. Likewise, it opens a whole new field of clinical research, such as the genetic factors associated to the susceptibility to this syndrome, the prognostic factors to present complications or to die, the effectiveness of the different treatments, among others.
Acknowledgements:
To the staff of the national reference laboratories for enteropathogens, acute respiratory infections, vector-borne viruses, respiratory viruses, and enteroviruses. To Dr. Juan Carlos Gomez de la Torre for his support in processing the gastrointestinal panels of 8 patients in 2018.
REFERENCES
1. Willison HJ, Jacobs BC, van Doorn PA. Guillain-Barré syndrome. Lancet. 2016;388(10045):717-727. doi: 10.1016/S0140-6736(16)00339-1. [ Links ]
2. Poropatich KO, Walker CLF, Black RE. Quantifying the association between Campylobacter infection and Guillain-Barré syndrome: a systematic review. J Health Popul Nutr. 2010;28(6):545-552. [ Links ]
3. McGrogan A, Madle GC, Seaman HE, de Vries CS. The epidemiology of Guillain-Barré syndrome worldwide. A systematic literature review. Neuroepidemiology. 2009;32(2):150-163. doi: 10.1159/000184748. [ Links ]
4. Martín Arias LH, Sanz R, Sáinz M, Treceño C, Carvajal A. Guillain-Barré syndrome and influenza vaccines: A meta-analysis. Vaccine. 2015;33(31):3773-3778. doi: 10.1016/j.vaccine.2015.05.013. [ Links ]
5. Musso D, Bossin H, Mallet HP, Besanrd M, Broult J, Baudouin L, et al. Zika virus in French Polynesia 2013-14: anatomy of a completed outbreak. Lancet Infect Dis. 2018;18(5):e172-e182. doi: 10.1016/S1473-3099(17)30446-2. [ Links ]
6. Villamil-Gomez WE, Sánchez-Herrera ÁR, Hernandez H, Hernández-Iriarte J, Díaz-Ricardo K, Castellanos J, et al. Guillain-Barré syndrome during the Zika virus outbreak in Sucre, Colombia, 2016. Travel Med Infect Dis. 2017;16:62-63. doi: 10.1016/j.tmaid.2017.03.012. [ Links ]
7. Dos Santos T, Rodriguez A, Almiron M, Sanhueza A, Ramon P, K de Oliveira W, et al. Zika Virus and the Guillain-Barré Syndrome - Case Series from Seven Countries. N Engl J Med. 2016;375(16):1598-1601. doi: 10.1056/NEJMc1609015. [ Links ]
8. Huang H-I, Shih S-R. Neurotropic Enterovirus Infections in the Central Nervous System. Viruses. 2015;7(11):6051-6066. doi: 10.3390/v7112920. [ Links ]
9. Messacar K, Abzug MJ, Dominguez SR. 2014 outbreak of enterovirus D68 in North America. J Med Virol. 2016;88(5):739-745. doi: 10.1002/jmv.24410. [ Links ]
10. Kim H, Kang B, Hwang S, Won Lee S, Cheon D-S, Kim K, et al. Clinical and enterovirus findings associated with acute flaccid paralysis in the Republic of Korea during the recent decade. J Med Virol. 2014;86(9):1584-1589. doi: 10.1002/jmv.23763. [ Links ]
11. Munayco CV, Soto Cabezas MG, Reyes MF, Arica Gutiérrez JA, Napanga Saldaña O. Epidemiología del síndrome de Guillain-Barré en el Perú. Rev Peru Med Exp Salud Pública. 2019;36(1):10. doi: 10.17843/rpmesp.2019.361.3729. [ Links ]
12. Dirección General de Epidemiología. Vigilancia, Prevención y Control de Síndrome de Guillan-Barré [Internet]. CDC-Perú; 2019 [citado el 2 de diciembre de 2019]. Disponible en: https://www.dge.gob.pe/portal/index.php?option=com_content&view=article&id=651&Itemid=418. [ Links ]
13. Rodríguez-Morales AJ, Failoc-Rojas VE, Díaz-Vélez C. Gastrointestinal, respiratory and/or arboviral infections? What is the cause of the Guillain-Barré syndrome epidemics in Perú? Current status - 2019. Travel Med Infect Dis. 2019;30:114-116. doi: 10.1016/j.tmaid.2019.06.015. [ Links ]
14. Ministerio de salud. Protocolo Sanitario de Urgencia para la vigilancia del síndrome de Guillain Barré, 2016 [Internet]. MINSA; 2016 [citado el 23 de octubre de 2019]. Disponible en: http://www.dge.gob.pe/portal/docs/tools/sg/psu13.PDF. [ Links ]
15. Ministerio de salud. Alerta Epidemiológica AE-CDC-No011-2019 -Incremento de casos de síndrome de Guillain Barré en varias regiones del país [Internet]. MINSA; 2019 [citado el 23 de octubre de 2019]. Disponible en: https://www.dge.gob.pe/portal/docs/alertas/2019/AE011.pdf. [ Links ]
16. Instituto Nacional de Salud. Laboratorios de Referencia Nacional [Internet]. INS [citado el 23 de octubre de 2019]. Disponible en: https://web.ins.gob.pe/es/salud-publica/enfermedades-transmisibles/laboratorios-de-referencia-nacional#vigilancia. [ Links ]
17. Totomoch-Serra A, Marquez MF, Cervantes-Barragán DE. Sanger sequencing as a first-line approach for molecular diagnosis of Andersen-Tawil syndrome. F1000Research. 2017; 6:1016. doi: 10.12688/f1000research.11610.1. [ Links ]
18. Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215(3):403-410. doi: 10.1016/S0022-2836(05)80360-2. [ Links ]
19. Hughes RAC, Rees JH. Clinical and Epidemiologic Features of Guillain-Barré Syndrome. J Infect Dis. 1997;176(s2):S92-S98. doi: 10.1086/513793. [ Links ]
20. Zhang M, Li Q, He L, Meng F, Zheng M, Gong Y, et al. Association Study Between an Outbreak of Guillain-Barre Syndrome in Jilin, China, and Preceding Campylobacter jejuni Infection. Foodborne Pathog Dis. 2010;7(8):913-919. doi:10.1089/fpd.2009.0493. [ Links ]
21. Platts-Mills JA, Liu J, Gratz J, Mguma E, Amour C, Swai N, et al. Detection of Campylobacter in Stool and Determination of Significance by Culture, Enzyme Immunoassay, and PCR in Developing Countries. J Clin Microbiol. 2014;52(4):1074-1080. doi: 10.1128/JCM.02935-13. [ Links ]
22. Kuwabara S, Yuki N. Axonal Guillain-Barré syndrome: concepts and controversies. Lancet Neurol. 2013;12(12):1180-1188. doi: 10.1016/S1474-4422(13)70215-1. [ Links ]
23. Jo Y-S, Choi J-Y, Chung H, Kim Y, Na S-J. Recurrent Guillain-Barré Syndrome Following Urinary Tract Infection by Escherichia coli. J Korean Med Sci. 2018;33(4):e29. doi: 10.3346/jkms.2018.33.e29. [ Links ]
24. Barbi L, Coelho AVC, Alencar LCA de, Crovella S. Prevalence of Guillain-Barré syndrome among Zika virus infected cases: a systematic review and meta-analysis. Braz J Infect Dis. 2018;22(2):137-141. doi: 10.1016/j.bjid.2018.02.005. [ Links ]
25. Agarwal A, Vibha D, Srivastava AK, Shukla G, Prasad K. Guillain-Barre syndrome complicating chikungunya virus infection. J Neurovirol. 2017;23(3):504-507. doi:10.1007/s13365-017-0516-1. [ Links ]
26. del Carpio-Orantes L, Da Silva IRF, Moguel KGP, Sánchez Díaz JS, Mata Miranda MDP, García-Méndez S, et al. Guillain Barré syndrome in arbovirus outbreak, Campylobacter claims his throne. J Neurol Sci. 2019; 396:254-255. doi: 10.1016/j.jns.2018.10.029. [ Links ]
27. Yamana M, Kuwahara M, Fukumoto Y, Yoshikawa K, Takada K, Kusunoki S. Guillain-Barré syndrome and related diseases after influenza virus infection. Neurol - Neuroimmunol Neuroinflammation. 2019;6(4):e575. doi: 10.1212/NXI.0000000000000575. [ Links ]
28. Kuroki S, Saida T, Nukina M, Haruta T, Yoshioka M, Kobayashi Y, et al. Campylobacter jejuni strains from patients with Guillain-Barré syndrome belong mostly to Penner serogroup 19 and contain beta-N-acetylglucosamine residues. Ann Neurol. 1993;33(3):243-247. doi: 10.1002/ana.410330304. [ Links ]
29. Mestanza F, Pamo O. Estudio muestral del consumo de medicamentos y automedicación en Lima Metropolitana. Rev Medica Hered. 1992;3(3). doi: 10.20453/rmh.v3i3.373. [ Links ]
30. Cruz N de la, Martin H. Automedicacion con antibioticos en estudiantes universitarios de Trujillo - Peru [tesis]. La Libertad: Facultad de Medicina Huamana, Universidad Privada Antenor Orrego; 2016. Disponible en: http://repositorio.upao.edu.pe/handle/upaorep/2125. [ Links ]
Received: January 27, 2020; Accepted: September 23, 2020










 text in
text in