Servicios Personalizados
Revista
Articulo
Indicadores
-
 Citado por SciELO
Citado por SciELO
Links relacionados
-
 Similares en
SciELO
Similares en
SciELO
Compartir
Revista Peruana de Medicina Experimental y Salud Publica
versión impresa ISSN 1726-4634
Rev. perú. med. exp. salud publica vol.37 no.4 Lima oct-dic 2020 Epub 11-Nov-2020
http://dx.doi.org/10.17843/rpmesp.2020.374.4825
Originales breves
Utility of flow citometry for detecting metallo-beta-lactamase-producing Pseudomonas aeruginosa
1 Hospital de Alta Complejidad Virgen de la Puerta, EsSalud, La Libertad, Perú.
2 Universidad Nacional de Trujillo, La Libertad, Perú.
4 Centro de Investigaciones Tecnológicas, Biomédicas y Medioambientales (CITBM), Universidad Nacional Mayor de San Marcos, Lima, Perú.
5 Laboratorio de Resistencia Bacteriana, Facultad de Farmacia y Bioquímica, Instituto de Investigaciones en Bacteriología y Virología Molecular (IBaViM), Universidad de Buenos Aires Argentina.
INTRODUCTION
Pseudomonas aeruginosa is one of the main opportunistic pathogens that cause a wide variety of healthcare-associated infections, such as sepsis, pneumonia, urinary tract, and soft tissue infections. In addition, it might be resistant, by intrinsic or acquired mechanisms, to several types of antibiotics, such as beta-lactams, aminoglycosides, fluoroquinolones and polymyxins. As a result, the treatment of P. aeruginosa infections is limited to certain classes of antibiotics, e. g. carbapenems, considered the most powerful antibiotic against this microorganism 1.
Carbapenems, such as imipenem (IPM) and meropenem (MEM), are the most effective antibiotics for treating infections caused by multidrug resistant (MDR) gram-negative bacteria, such as P. aeruginosa 2. Carbapenem resistance can be divided into two types: non-carbapenemase-producing (Non-CP) mechanisms and carbapenemase-producing (CP) mechanisms. Non-CP mechanisms have lower sensitivity to carbapenems due to the loss of porins (OprD) or the overexpression of AmpC beta-lactamases and efflux pumps 3. In contrast, the CP mechanism acquires a transmissible gene that produces a specific enzyme for the hydrolysis of carbapenems, a carbapenemase 4.
The production of carbapenemases is one of the most important mechanisms in which P. aeruginosa becomes resistant to carbapenems by mobile genetic elements. These enzymes are beta-lactamases, which hydrolyze carbapenems, as well as almost all beta-lactams. Based on Ambler’s molecular classification, carbapenemases are divided into class A, serine-beta-lactamases (SBL); class B, metallo-beta-lactamases (MBL); and classes C and D, also SBL. While SBL requires a serine residue in its active site, MBL requires divalent metal cations, such as zinc for enzymatic activity 1 , 5.
During clinical practice, rapid and accurate detection of CP P. aeruginosa is considered essential for implementing timely and specific treatment for the infection 9. However, this process takes an average of 55 hours between the sample collection for the culture and the delivery of the antibiogram report, and mortality increases by 7.6% for each hour of delay in the administration of an effective treatment 6.
In Peru, the frequency of MBL in carbapenem-resistant P. aeruginosa has increased in the last decade, from 15.7% in 2011 7 to 31.6% in 2016 8, evidencing that this resistance mechanism is present in our environment; therefore, it is important to ensure a rapid and accurate detection by clinical microbiology laboratories.
Several phenotypic tests based on inhibitors have been proposed for screening MBL-producing P. aeruginosa, however, all of them must be confirmed by molecular biology techniques for gene identification. Phenotypic methods use metal chelators such as ethylenediaminetetraacetic acid (EDTA) or dipicolinic acid (DPA) 9.
Flow cytometry (FC) is a quantitative analysis technology used to characterize cell populations individually. Cells are illuminated by a laser lamp, then the obtained signal intensities are detected by a fluorescence sensor and are finally correlated with structural or functional parameters of the analyzed cell for classification 10. The aim of this study was to evaluate the usefulness of FC for the detection of MBL-producing P. aeruginosa.
KEY MESSAGES
Motivation for the study: Pseudomonas aeruginosa is an opportunistic bacterium that causes healthcare-associated infections where resistance to carbapenems is increasing. We present a flow cytometry (FC) method for detecting metallo-beta-lactamases (MBL).
Main findings: With the FC method for detecting MBL, we obtained a sensitivity of 94.1% and a specificity of 100%, as well as a positive predictive value of 100%.
Implications: FC assays could provide a rapid and intuitive method for detecting MBL-producing P. aeruginosa.
THE STUDY
A descriptive study of isolates of Pseudomonas aeruginosa resistant to carbapenems identified genotypically as MBL carriers (17 strains: 14 with the blaIMP gene and 3 with the blaVIM gene) and not MBL carriers (12 strains), collected between 2011 and 2017 from the strain collection of the Laboratory of Molecular Epidemiology and Genetics of the Instituto de Medicina Tropical Daniel A. Carrión of Universidad Nacional Mayor de San Marcos. The isolates were then transported to the Clinical Pathology Laboratory of Hospital de Alta Complejidad Virgen de la Puerta, EsSalud, where they were stored at -20 °C until they were processed.
The strains were reactivated to evaluate their purity and viability. Then, we made two bacterial suspensions (0.5 McFarland scale) by isolation. One bacterial suspension was incubated with MEM PHARMAGEN® (8 µg/mL); the other bacterial suspension was incubated with MEM PHARMAGEN® (8 µg/mL) plus EDTA Merk® inhibitor (12.5 mM). Both suspensions were incubated at 37 °C for two hours 11.
After two hours of incubation, the viability test was carried out with the Becton Dickinson (BD) cell viability kit. The cell viability kit reagent (5 µL of Thiazole Orange [TO], 42 µmol/L in dimethyl sulfoxide and 5 µL of propidium iodide [PI] 4.3 mmol/L in water) was added to 500 µL of the previously treated bacterial suspension. Then, the suspension was mixed and incubated for 5 minutes at room temperature in a dark room. Voltages were adjusted graphically to differentiate the bacterial population marked with TO, a cellularity marker, from debris.
The reading was performed in the FACSCalibur BD flow cytometer. The results were interpreted according to the increase or not of fluorescence emitted by the PI, a positive MBL detection test was considered when observed a significant increase in the fluorescence ratio of the suspension treated with MEM plus EDTA compared to the MEM only incubated suspension. A control tube was used during the whole procedure, where only the bacterial suspension of each strain was incubated without any antibiotic or inhibitor, and was treated with the cellular viability (BD) kit to confirm the viability of the bacteria against the reagents used.
The samples were analyzed with the Cell Quest® program, where bacteria were classified according to the fluorescence emitted, quantifying it as an “event”. If an event was positive only for TO, it was classified as “alive”; if it was positive for PI and TO, it was classified as “damaged”, and if it was positive only for PI, it was classified as “degraded”. To obtain a measurable and comparable quantity, the rate of increase events obtained between the tube with MEM-EDTA and the tube with MEM (% tube events with MEM-EDTA / % tube events with MEM) was compared with the percentage of events obtained, in the MBL-producing isolates and the non-MBL-producing isolates.
We carried out the student’s t-test, the analysis of the ROC curve, and determined sensitivity and specificity. We used SPSS 24 statistical program.
The Ethics Committee of Universidad Nacional de Trujillo approved this study’s protocol. All the procedures in this study were carried out in accordance with the good clinical practice and ethics in biomedical research guidelines.
FINDINGS
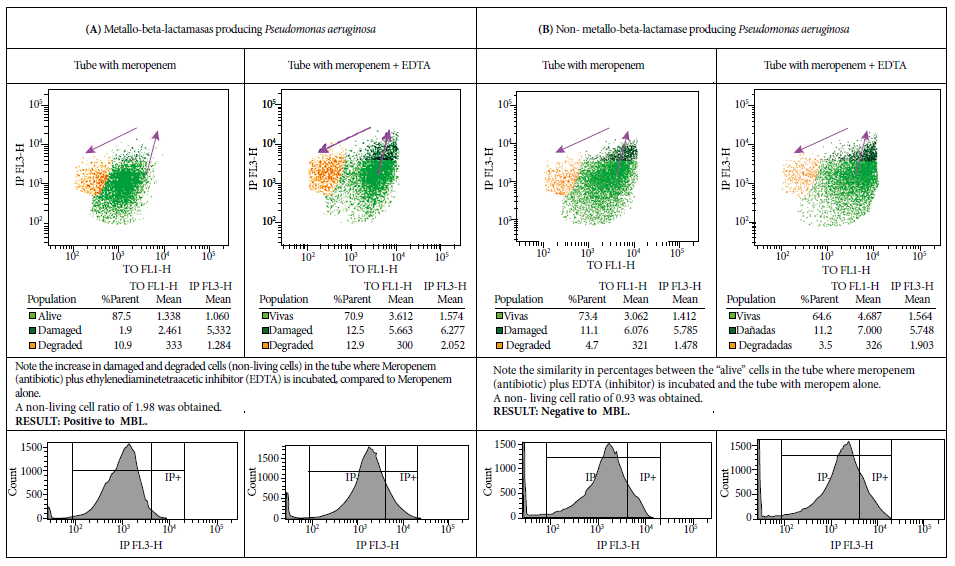
With FC and cell viability kit, we identified 16 MBL-producing P. aeruginosa and 13 non-MBL producing P. aeruginosa. When comparing the fluorescence increase index (the sum of the percentage of events presented in cells classified as “damaged” and “degraded”) among “non-living” cells in the tubes with antibiotic plus inhibitor with those in the tube with only antibiotic, we obtained an average index of 3.17 for MBL-producing P. aeruginosa, which showed a significant difference (p<0.01) (Figure 1).
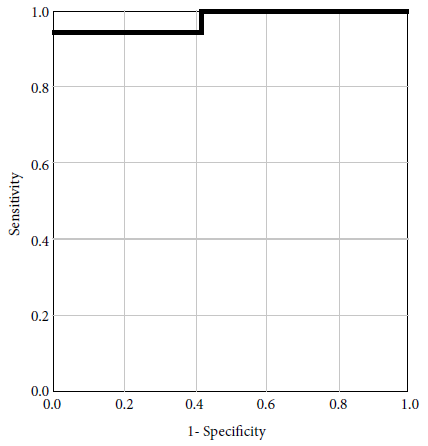
To evaluate the sensitivity and specificity of FC for MBL detection, we carried out a ROC curve analysis and found an area under the curve (AUC) of 0.975 (Figure 1). This suggests a best cut off point of 1.6 for the “non-living” cell index.
Using the genotypic results (gold standard) and the results obtained by flow cytometry, as well as the non-living” cells index cut-off point of 1.6, we obtained a sensitivity of 94.1% and a specificity of 100%. We also obtained a positive predictive value (PPV) of 100% (Figure 2).
DISCUSSION
We introduce an alternative method for detecting MBL in P. aeruginosa using FC and BD cell viability kit, by evaluating the bacterial strain alone, the strain against the carbapenem antibiotic (MEM) and the strain against the antibiotic plus MBL inhibitor (EDTA). We obtained satisfactory results of 94.1% sensitivity and 100% specificity.
Mulroney et al. 12 and Kilic et al. 13 have used FC for just two hours as a predictor of resistance to carbapenems. However, they did not establish the presence or absence of carbapenemases. Pina-Vaz et al. 14 combines FC with computer analysis for detecting classes A, B, and D carbapenemases in a small number of isolates, without reporting sensitivity or specificity. Silva et al. 11 used FC for detecting carbapenemases in enterobacteria (30 CP and 30 non-CP) and reported 100% sensitivity and specificity. These results are similar to ours. The difference in sensitivity could be a result of our work with a different bacterial group, and since only one strain (1/17 MBL) of P. aeruginosa carrying the blaIMP gene was not detected by FC, we considered it as a false negative; this is possibly due to a low expression of this type of MBL enzymes in the studied strain.
Other studies have used FC for detecting beta-lactamases, like the one carried out by Faria-Ramos et al. 15, where they used the FACSCalibur FC to search for extended spectrum beta-lactamases (ESBL) in enterobacteria, with a fluorescence increase index of 1.5 between the antibiotic plus inhibitor (clavulanic acid) and the antibiotic alone, similar to the index reported in our research. On the other hand, Akhmaltdinova 16 used microdilution as a reference and compared it to FC regarding BLEE detection and reported 85.7% sensitivity and 88.8% specificity.
Within the phenotypic tests for MBL detection, Heinrichs et al. 7 compared different phenotypic methods in 183 P. aeruginosa isolates, using diffusion discs with imipenem (10 µg) + EDTA (750 µg), and reported a sensitivity of 90% and a specificity of 93%; with meropenem (10 µg) + DPA (1000 µg) they obtained a sensitivity of 97% and specificity of 88%. Looking for synergy, with DPA + imipenem 10 µg / meropenem 10 µg they obtained a sensitivity and specificity of 100% and 31%, respectively. When they evaluated the sensitivity and specificity of colorimetric methods, such as CarbaNP®, they found 88% and 99%, respectively.
Therefore, FC is more advantageous than culture-based methods that must wait 24 hours for interpretation. FC can provide us with a preliminary result in two hours while the confirmation happens in the next 24 hours. Another important point is that CF avoids subjective interpretation when observing color change, which happens with colorimetric rapid tests, making it a tool with which we can achieve an objective analysis by estimating fluorescence using indices, while having a CF team could easily become a guidance tool for the search of MBL in P. aeruginosa in a short time.
One of the limitations of this study is the limited number of samples, because we included only confirmed genotypic isolates. Future research on multiple mechanisms of resistance and simultaneous expression (serin carbapenemases and ESBL) could determine the specificity for MBL using this study for P. aeruginosa, although we must consider that MBL production is frequent in this microorganism.
In conclusion, our results suggest that FC assays, considering cell viability with antibiotic and antibiotic plus inhibitor, could provide a quick and intuitive method to detect presumptive MBL-producing P. aeruginosa. However, further studies with larger numbers of isolates are needed to confirm the accuracy of this methodology.
Acknowledgements:
To the Laboratory personnel of Hospital de Alta Complejidad Virgen de la Puerta for their support, to the Laboratory of Molecular Epidemiology and Genetics of Instituto de Medicina Tropical Daniel A. Carrión UNMSM for facilitating the strains for the realization of the study, to Dr. Patricia Contreras and to Mg. Paul Ríos Sanca for their collaboration in the execution of this research.
REFERENCES
1. Hong DJ, Bae IK, Jang IH, Jeong SH, Kang HK, Lee K. Epidemiology and characteristics of Metallo-ß-Lactamase-producing Pseudomonas aeruginosa. Infect Chemother. 2015;47(2):81-97. doi: 10.3947/ic.2015.47.2.81. [ Links ]
2. He Q, Chen W, Huang L, Lin Q, Zhang J, Lui R, et al. Perfomance evaluation of three automated identification systems in detecting carbapenem-resistant Enterobacteriaceae. Ann Clin Microbiol Antimicrob. 2016;15(1):40. doi: 10.1186/s12941-016-0154-0. [ Links ]
3. Nordmann P, Poirel L, Dortet L. Rapid detection of carbapenemse-producing Enterobacteriaceae. Emerg Infect Dis. 2012;18(9):1503-7. doi: 10.3201/eid1809.120355. [ Links ]
4. Banerjee R, Humphries R. Clinical and laboratory considerations for rapid detection of carbapenem-resistant Enterobacteriaceae. Virulence. 2017;8(4):427-439. doi: 10.1080/21505594.2016.1185577. [ Links ]
5. Sacha P, Wieczorek P, Hauschild T, Zórawski M, Olszanska D, Tryniszewska E. Metallo-ß-lactamases of Pseudomonas aeruginosa a novel mechanism resistance to ß-lactam antibiotics. Folia Histochem Cytobiol. 2008;46(2):137-42. doi: 10.2478/v10042-008-0020-9. [ Links ]
6. Kumar MS, Lakshmi V, Rajagopalan R. Ocurrence of extended spectrum ß-lactamases among Enterobacteriaceae spp. Isolated at a tertiary care institute. Indian J Med Microbiol. 2006;24(3):208-11. [ Links ]
7. Gonzales-Escalante E, Vicente-Taboada W, Champi-Merino R, Soto-Pastrana J, Flores-Paredes W, Lovera-García M, et al. Metalo-ß-lactamasas en aislamientos clínicos de Pseudomonas aeruginosa en Lima, Perú. Rev Peru Med Exp Salud Publica. 2013;30(2):241-5. [ Links ]
8. Salvador-Luján G, García-de-la-Guarda R, Gonzales-Escalante E. Caracterización de metalo-ß-lactamasas en aislados clínicos de Pseudomonas aeruginosa recuperados de pacientes hospitalizados en el Hospital Militar Central. Rev Peru Med Exp Salud Publica. 2018;35(4):636-641. doi: 10.17843/rpmesp.2018.354.3755. [ Links ]
9. Heinrichs A, Huang TD, Berhin C, Boagerts P, Glupczynski Y. Evaluation of several phenotypic methods for the detection of carbapenemse-producing Pseudomonas aeruginosa. Eur J Clin Microbiol Infect Dis. 2015;34(7):1467-74. doi: 10.1007/s10096-015-2376-z. [ Links ]
10. Díaz M, Herrero M, García AL, Quirós C. Application of flow cytometry to industrial microbial bioprocesses. Biochem Eng J. 2010; 48(3):385-407. doi: 10.1016/j.bej.2009.07.013. [ Links ]
11. Silva A, Faria-Ramos I, Ricardo E, Miranda MI, Espinar MJ, Costa-de-Oliveira S, et al. Rapid flow cytometry test for identification of different carbapenemses in Enterobacteriaceae. Antimicrob Agents Chemother. 2016;60(6):3824-6. doi: 10.1128/AAC.02947-15. [ Links ]
12. Mulroney KT, Hall JM, Huang X, Turnbull E, Bzdyl NM, Chakera A, et al. Rapid susceptibility profiling of carbapenem-resistant Klebsiella pneumoniae. Sci Rep. 2017;7(1):1903. doi: 10.1038/s41598-017-02009-3. [ Links ]
13. Kilic A, Dogan E, Kaya S, Oren S, Tok D, Ardic N, et al. Rapid Identification of Klebsiella pneumoniae by Matrix-Assisted Laser Desorption/Ionization-Time of Flight Mass Spectrometry and Detection of Meropenem Resistance by Flow Cytometric Assay. J Clin Lab Anal. 2016;30(6):1191-1197. doi: 10.1002/jcla.22002. [ Links ]
14. Pina-Vaz C, Silva AP, Faria-Ramos I, Teixeira-Santos R, Moura D, Vieira TF, et al. A Flow Cytometric and Computational Approaches to Carbapenems Affinity to the Different Types of Carbapenemases. Front Microbiol. 2016; 7: 1259. doi: 10.3389/fmicb.2016.01259. [ Links ]
15. Faria-Ramos I, Espinar MJ, Rocha R, Santos-Antunes J, Rodrigues AG, Cantón R, et al. A novel flow cytometric assay for rapid detection of extended-spectrum ß-lactamases. Clin Microbiol Infect. 2013;19(1):E8-E15. doi: 10.1111/j.1469-0691.2012.03986.x. [ Links ]
16. Akhmaltdinova L, Lavrinenko A, Belyayev I. Flow Cytometry in Detecting Resistant E. coli Strains. Open Access Maced J Med Sci. 2017 Jul 28;5(5):592-594. doi: 10.3889/oamjms.2017.104. [ Links ]
Cite as: Salinas-Salcedo IA, Saldaña-Jiménez JA, Guevara-Canales JM, Gonzales-Escalante E. Utility of flow cytometry for detecting metallo-beta-lactamase-producing Pseudomonas aeruginosa. Rev Peru Med Exp Salud Publica. 2020;37(3). doi: https://doi.org/10.17843/rpmesp.2020.374.4825.
Received: September 22, 2019; Accepted: July 16, 2020










 texto en
texto en